Nucleosome Structure Part 1
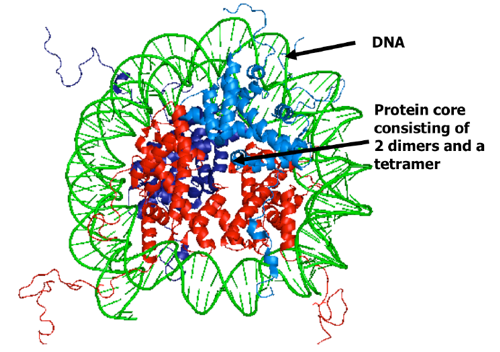
Nucleosome Structure - Part 1
Proteins, Nucleic Acids, and other molecules can form crystals in appropriate conditions which are repeating “unit cells” that has a specific and uniform pattern. The nucleosome (DNA wrapped twice around 8 histones) can also be crystallized and analyzed (see the picture).
Structure of the Nucleosome Crystal:
The structure of the histone was determined in the late 1990’s so it’s a pretty recent discovery. It consists of 147 Base Pairs of DNA (Adenine-Thymine, Guanine-Cytosine) are wrapped around 8 histones (2 sets of H2A, H2B, H3, H4 histones).
The Folding of a single Histone:
A histone has a simple fold. It consists of a 4 sites with the same core. These 4 sites are: HSH1, C, N, HSH2 (see picture). The identical cores of the 4 types of histones suggests that they shared the same evolutionary parent. There is a histone protecting every 10 or 11 base pairs and exposes 1 phosphodiester bond which is why DNAse I cut fragments are cut into a minimum of 10 or 11 bases (Read my other article for more on DNAse I).
What composes the histone octomer (8 monomers joined)
The H3 and H4 histones are monomers individually like the H2A and H2B histones. Joined together, the pair forms the H3-H4 dimer (composed of 2 monomers). The H3-H4 dimer pairs with another H3-H4 dimer to form the H3-H4 Tetramer (4 monomers or 2 dimers). The H2A and H2B histones are joined the same with. Two monomer histones H2A and H2B jon to make the H2A-H2b dimer. This dimer joins with another of the same dimer to make the H2A-H2B tetramer. Both the H3-H4 Tetramer and H2A-H2B tetramer are like half of an orange. The two halves join to make the histone octomer (which is the unit that DNA wraps around twice).
How is DNA bent around the histone?
The histones are not prefect spheres. Actually, they look more like a sea urchin with spikes that do not form a perfect sphere. DNA is thus not bent smoothly on the histone. A gyre is a single wrapping of DNA around a histone. Because the histone isn’t a perfect spiky sphere and because the shape of DNA isn’t a perfectly straight helix, the DNA is bent with the length of the groove ranging from 0.5 Angstroms – 6.5 Angstroms of increasing length as the DNA twists along the histone (0.5A, 1.5A,, 2.5A, 3.5A, 4.5A, 5.5A, and 6.5A).
Role of Water on the histone and DNA
Simply put, water mediates the attachment between the phosphates on the DNA backbone (alternating sugar and phosphates) and the histone. Water is like the DNA Packaging Glue.
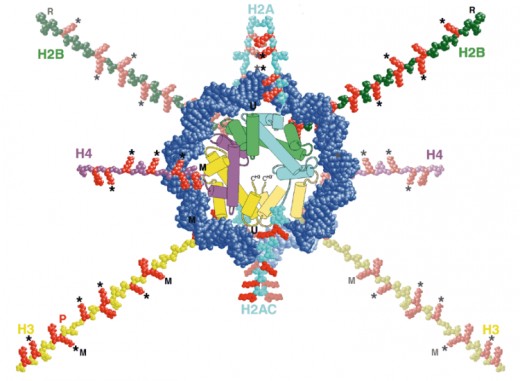
Histones have tails that stick out between the two DNA coils around it
Waving out of the histone core and in between the two coils of DNA is the histone tails that look similar to curly hair. These tails have been modified to control function. It is important in sequence reading and tells what kind of DNA the sequence is (since the sequences of the compressed DNA can’t be accessed by other reading proteins) whether a sequence of DNA is a gene, an intron (non-coding DNA), exon, etc.
Ways the Histone tails can be modified
Each histone modification determines a different function. Histones can be modified by: Acetylation, Methylation, Phosphorylation, and Ubiquitylation. Different combinations of modifications tell what different DNA sequences are. These marks are thus called “Epigenetic Marks” because they control gene function rather than be controlled by it (and that includes the packaging of DNA). They are independent from gene control. The combination of tail modifications are called the “histone code”
Histone Code Function
The histone code is epigenetic and so they work independently from DNA control but like DNA they are also inherited from both the parents and mashed together to form a different combination (unlike DNA which is exactly the pairing of chromosomes from both parents). The specific functions of these markers are not yet well understood. A few of the simpler functions have been noted, but not the complex ones.
Some of the combinations that have some understanding are:
Markers on Histone H3:
No Modifications – Gene silencing
Acetylated N-Terminal 14 – gene expression
Acetylated N-Terminal 9 – Histone deposition
Methylated N-Terminal 9 – Gene silencing/heterochromatin
Phosphorylated N-Terminal 10 AND Phosphorylated N-Terminal 28 – Mitosis/Meiosis
Phosphorylated N-Terminal 10 AND Acetylated N-Terminal 14 – Gene expression
Markers on Histone H4:
No Modifications – Gene silencing
Acetylated N-Terminal 5 and Acetylated N-Terminal 12 – Histone deposition
Acetylated N-Terminal 8 and Acetylated N-Terminal 16 – Gene expression