Unknown Determination
Unknown Organism Determination
The following is my official diagnostic report detailing my experimentation methodology in determining an "unknown" organism given for a course assignment in Microbiology (Fall semester 2010).
Unknown Organism Determination
Introduction:
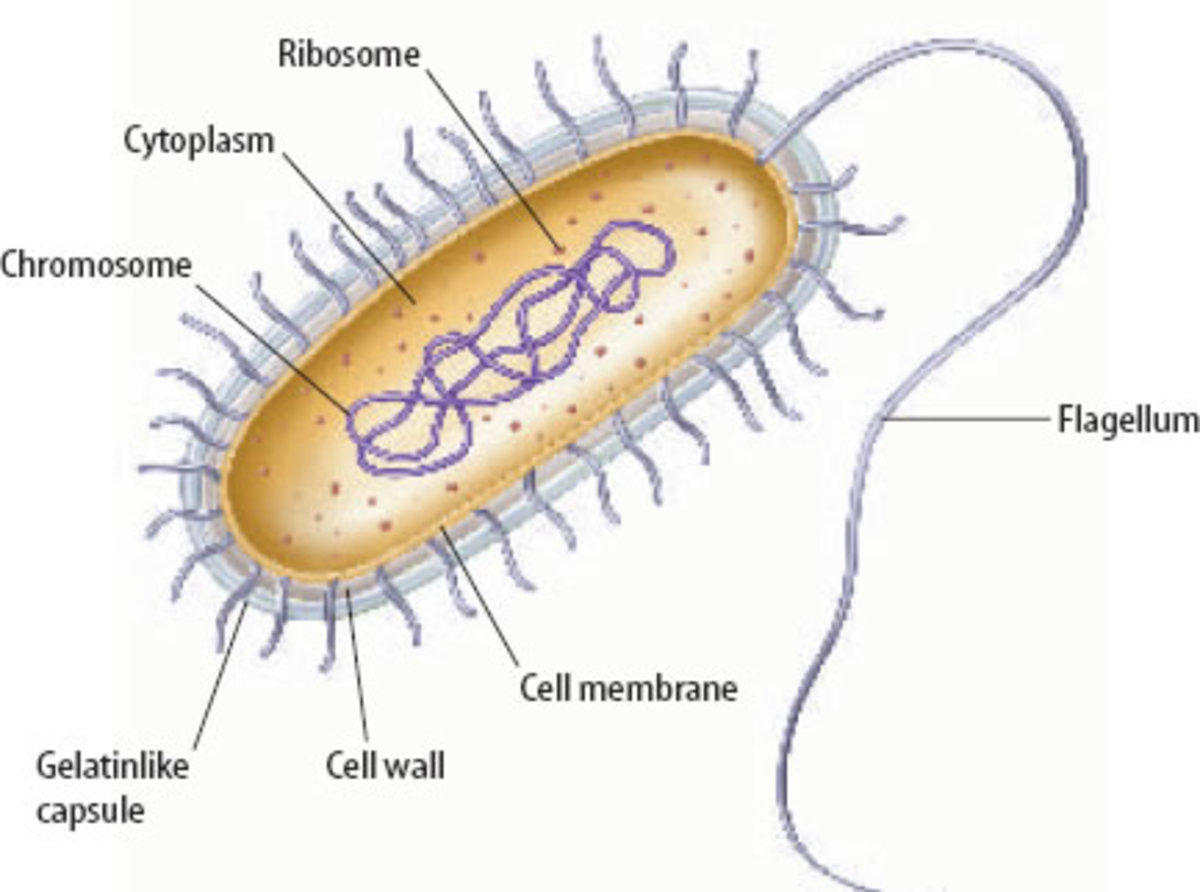
Enterococcus faecalis is a commensal, non-motile, facultative anaerobic bacterium, often found in fecal flora. It is classified as an enteric (intestinal) Enterococcus species, and traditionally classified as a part of the Group D Streptococcus genus of microorganisms. It appears as Gram-positive cocci in pairs or chains under the microscope and is found living in the gastro-intestinal tract at high salt concentrations in bile. E. faecalis can cause life-threatening infections due to multidrug antibiotic resistance and virulence factors, especially in nosocomial environments. Enterococci have been isolated from food, soil, plants and water and are often transferred from person to person through physical contact. It can survive harsh conditions in nature such as extreme temperatures and bile salts. When ingested, the pathogenic properties of the organism can cause infections of the bladder, urinary tract, prostate, epididymis, nervous system, surgical wounds and endocarditis. The steps repeated as documented, instructs the scientific procedure to be followed in the accurate diagnosis of organisms within the Enterococcus species as performed by myself.
Materials and Methods:
Materials used include: organism, gloves, flint striker, Bunsen burner, 2 microscope slides, microscope, inoculating loop, inoculating needle, test tube rack, water bottle, wax pencil, slide tray, Bile Esculin Azide Agar, tube of 6.5% NaCl broth, Phenol Red sucrose fermentation broth with Durham tube, Phenol Red mannitol fermentation broth with Durham tube, BHI (brain heart infusion) agar plate, motility medium.
Methods: First, I prepared two heat-fixed smears which involves dropping a loop full of water onto a clean slide using an inoculating loop, transferring the culture to the water using aseptic technique and then heat fix by passing the smear through the top part of the inner blue cone of the Bunsen burner flame 3-4 times. This protocol kills the cells, causes the cells to stick to the slide and retain the dyes. I made sure to air dry the smear completely before running it through the Bunsen burner flame so as not to create living aerosols. I Gram-stained one of them and Simple stained the other. To prepare a Simple stain, I added a few drops of crystal-violet dye to one of my heat smears and let it sit for 60 seconds before I rinsed it with water and blotted dry. I prepared a Gram stain with the other heat smear which involves a few extra steps. After the crystal-violet step, I rinsed with water, added Iodine, rinsed with water again, decolorized with an alcohol mixture, rinsed with water to stop decolorization, added Safranin dye, rinsed with water and blotted dry. The Safranin dye is a negative dye which stains the positive components of the cells a “pink” color. I observed the Gram-stained slide under a microscope at 1000x magnification and discovered only violet-stained cocci in chained arrangements. No “pink” was observed in the stain, indicating that only the negative components of the organism were stained with the positively charged crystal-violet dye, which confirms the organism to be Gram-positive and not Gram-negative. This means that I was working with either a Streptococcus or Enterococcus species. Next, I performed a KOH test to further confirm that the organism was a Gram-positive species and to make sure I had not misinterpreted what I had seen under the microscope. For the KOH test, I added 2-3 drops of 10% potassium hydroxide (KOH) to a small drop of distilled water onto a clean microscope slide, transferred a visible clump of organism to the KOH solution using my inoculating loop, mixed the cells into the solution using small, circular motions for 60 seconds and then periodically lifted up the loop to look for what appears to be a stretchy mucous known as “stringing,” using aseptic technique. With the KOH test, no evidence of “stringing” was observed which confirms my observations under the microscope. Then I would continue on to perform the next steps which were to inoculate a tube of Bile Esculin Azide Agar and a tube of 6.5% NaCl broth with the unknown organism to determine whether I had an oral Streptococcus species or an intestinal Enterococcus species. The sodium azide in the Bile Esculin Azide Agar inhibits Gram-negative enterics while the bile inhibits Gram-positive organisms except the group D Streptococcus (Enterococcus). The test is performed “to see if the organism can hydrolyze esculin in the presence of bile” which will differentiate oral Streptococcus from Enterococcus. After the Bile Esculin Hydrolysis Test, I performed a test using a broth medium containing 6.5% NaCl, which will select for Enterococcus species that can withstand the high salt concentration, unlike Streptococcus species of the oral cavity. I aseptically transferred the organism into the 6.5% NaCl broth and incubated both tubes at 35 degrees Celsius for 48 hours. The next day, I interpreted and recorded the results from the previous day’s experimentation and then performed the rest of my experiments. I was able to determine from these results that I had an Enterococcus species and would need to do a Phenol Red carbohydrate fermentation test using sucrose to differentiate E. durans from other Enterococcus species, a Phenol Red carbohydrate fermentation test using mannitol to differentiate E. hirae from other Enterococcus species, and a BHI and motility test to differentiate E. faecium, E. faecalis and E. gallinarum. I decided to do all these tests at once to get a quicker result in a shorter time period. So, I transferred my organism to the tube of Phenol Red sucrose fermentation broth, the tube of Phenol Red mannitol fermentation broth, and swirled the organism around in the broth, knocking the organism into the medium, maintaining aseptic technique. Next, I transferred a loop full of the organism to the surface of a BHI agar plate using aseptic technique, and stored the plate upside-down with the lid on the bottom so that condensation that may occur during the incubation process would not fall onto the colonies, scattering the organism colonies. Then, I slowly stab-inoculated the organism into a motility medium which is “colorless in its oxidized form but turns red when reduced by microorganisms,” using aseptic technique. Last, I incubated these tubes and plate at 37 degrees Celcius for 48 hours. The next day I would record and interpret my results which would determine what species of Enterococcus I was dealing with.
Results:
KOH Test - Negative - No stringing
Bile Esculin Azide Agar Test - Positive - Darkening of Medium
6.5% NaCl Test - Positive - Turbidity and Sediment observed
Phenol Red Sucrose Fermentation - Positive - Red turned Yellow (positive for acid production)
Phenol Red Mannitol Fermentation - Positive - Red turned Yellow (positive for acid production)
BHI Test - Positive - Growth on Medium
Motility - Negative - No movement from stab line
Discussion/Conclusion:
The concluding results in my experimentation yielded a clear result for Enterococcus faecalis after careful analyzation of the prepared slides. As I mentioned earlier, the KOH test indicated no stringing which is a characteristic of only Gram-positive organisms. Then, looking at the results for the Bile Esculin Hydrolysis Test, I observed that the “yellow” colored medium had darkened to a brownish black, indicating that esculin was hydrolyzed and that the organism was an Enterococcus species. After incubation of the organism in the 6.5% NaCl broth, I examined the tube for evidence of growth in which I noticed slight turbidity and a notable amount of sediment considered to be growth. Since the organism was bile tolerant and able to grow in such a high salt concentration, I determined that the organism was an intestinal Enterococcus species once again. Following differentiation of the Enterococcus species from the Streptococcus species, further differentiation within the genre was needed to determine the unknown organism from E. durans, E. hirae, E. faecium, E. faecalis, and E. gallinarum. The next day I was able to examine results for the Phenol Red carbohydrate fermentation tubes, BHI and motility tests. Since Phenol Red is a pH indicator that turns yellow under acidic conditions, I concluded that the sucrose and mannitol fermentation tubes were positive for acid production because they had turned “yellow.” Durham tubes inside the Phenol Red carbohydrate fermentation tubes were also examined for the presence of gas. Acid production was present in the Phenol Red sucrose fermentation tube which ruled out a member of the Enterococcus species known as E. durans and a positive result for acid production in the Phenol Red mannitol fermentation tube ruled out a member of the Enterococcus species known as E. hirae. Growth on the BHI agar plate indicated that the organism could not be E. faecium because it cannot grow on BHI agar plates supplemented with 0.04% potassium tellurite which behaves as an antibiotic. Last, I examined the motility tube for motility and observed that the organism had not migrated away from the stab line, which ruled out E. gallinarum; an Enterococcus species that is positive for motility. Therefore, since the unknown organism was non-motile and grew on the BHI agar plate, I came to the final resolution that my organism was Enterococcus faecalis.
References:
1. Carrington, Elizabeth, Ph.D. Microbiology Laboratory Manual. Second Edition. Dubuque: Kendall Hunt Publishing Company, 2006, 2009.
2. Fischetti, Vincent A. Gram-Positive Pathogens. Washington D.C.: ASM Press, 2006.
3. Gilmore, Michael S. The Enterococci: Pathogenesis, Molecular Biology, and Antibiotic Resistance. Washington D.C.: ASM Press, 2002.
4. Murray, Barbara E. Emerging Infectious Diseases. Mar. 1998. Centers for Disease Control and Prevention. <http://www.cdc.gov/ncidod/eid/vol4no1/adobe/murray.pdf>.
5. Pelletier LL (1996). Microbiology of the Circulatory System. In: Baron's Medical Microbiology (Baron S et al., eds.) (4th ed.). Univ of Texas Medical Branch.
6. Robinson, D. Ashley, Daniel Falush and Edward J. Feil. Bacterial Population Genetics in Infectious Disease. Hoboken: Wiley & Sons, 2010.
7. Toy, Eugene C. and Audrey Wanger. Case Files: Microbiology, Second Edition. The McGraw-Hill Companies, Inc. 2005.
8. Wood, Brian J. B. and W. H. Holzapfel. The Lactic Acid Bacteria: The Genera of Lactic Acid Bacteria. New York: Chapman & Hall, 1995.