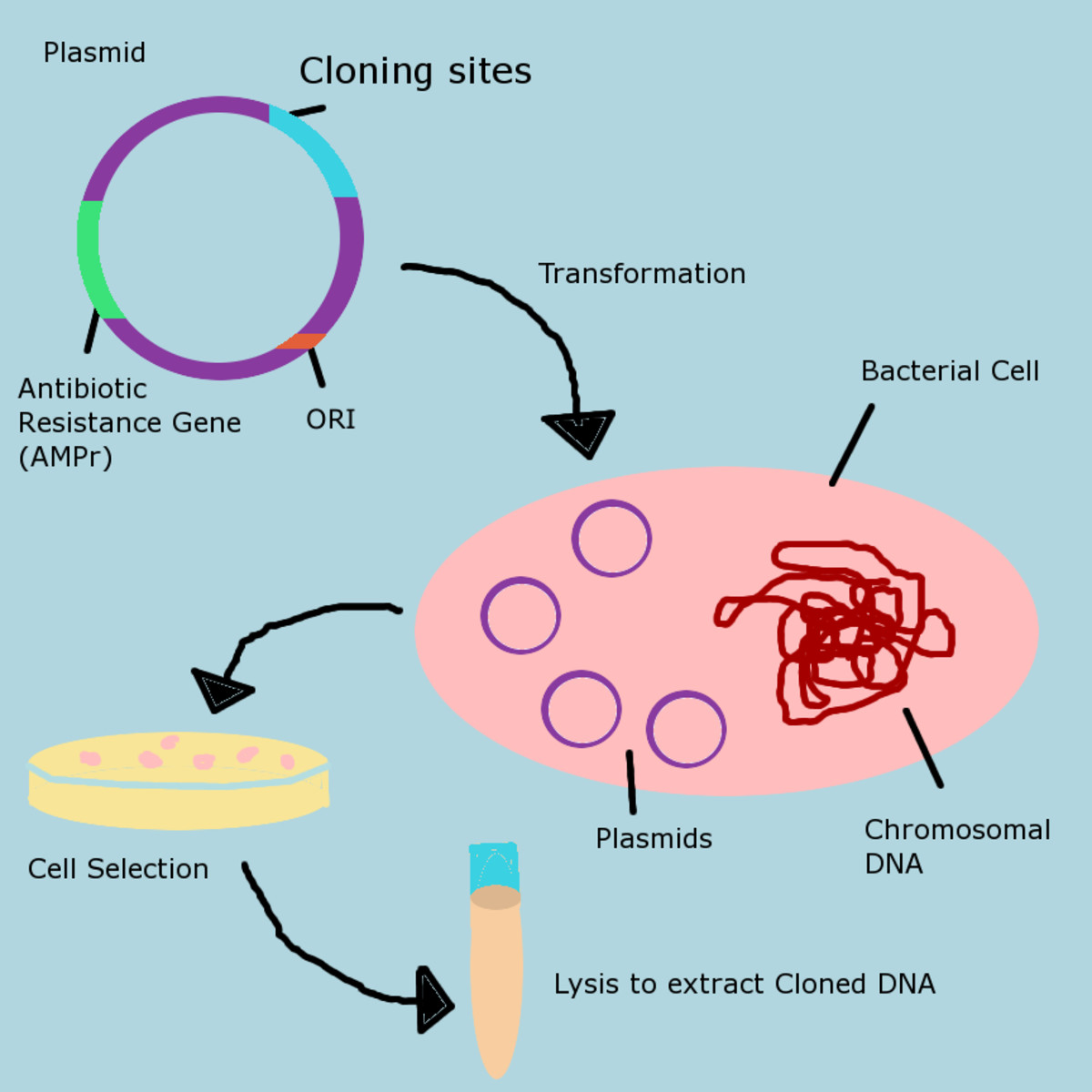
Genetic Engineering Basics
1. Obtaining the gene to be engineered
cells in the islets of langerhan in the pancreas. This mRNA can then be used as a template to produce a copy of the required gene.
Another process of obtaining a certain gene (and the process I will focus on in this hub) is using a DNA probe to identify the location of the gene which is then is cut from the DNA using restriction enzymes. The basic process of obtaining the gene in this way is outlined in the following steps:
- Once the gene has been located, specific restriction enzymes are used to cut through the DNA at specific points where a specific base sequence occurs - the restriction site.
- The enzyme catalyses a hydrolysis reaction which breaks the sugar-phosphate backbone of the DNA at the specific points which leaves a 'staggered cut' with exposed bases, these are known as 'sticky ends'.
2. A copy of the gene is placed in a vector
After the gene has been cut from the DNA using restriction enzymes it is then placed in a vector (the agent that carries a section of DNA from one cell to another). The majority of vectors used are bacterial plasmids which is a circular section of DNA that are found in many types of bacteria and are separate from the main bacterial chromosome. The plasmids often contain antibiotic resistant genes which are used as genetic markers later on in the genetic engineering process. However, some other vectors such as yeast chromosomes and virus genomes are used as a vector.
The plasmids are cut with the same restriction enzymes that were used to isolate the gene needed to be engineered - this ensures that the sticky ends are complementary and thus the gene and the plasmid can join. The cut plasmids, the isolated gene and ligase enzymes are then mixed together and some of the plasmids will take up the gene and ligase enzymes will catalyse the joining of the sugar-phosphate backbone of the gene and the plasmid in a condensation reaction. This forms a recombinant plasmid - meaning a plasmid that has genetic material from two different organisms. However, not all of the plasmids will combine with the gene - many plasmids will simply reseal to form the original plasmid (with the help of ligase!).
Large quantities of the plasmid and bacteria cells are then mixed in the presence of calcium salts, the temperature is also reduced to freezing and then heated to around 40 degrees, this ensures the maximum amount of bacterial cells take up the plasmid. The bacteria cells that do take up the recombinant plasmid are then known as 'transformed' bacterial cells and are by definition transgenic.
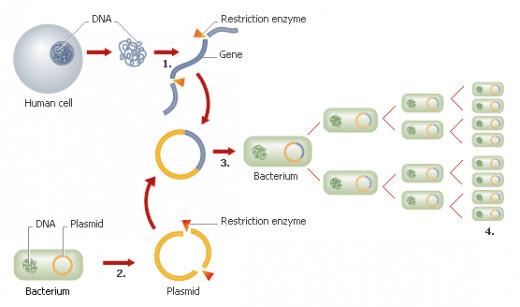
3. The vector carries the gene to the recipient cell
There are a number of ways to get the vector containing the gene into the recipient cell, some of which include:
- Microinjection - small sections of the DNA can be directly injected into the recipients cell nucleus using a very small micropipette.
- Electroporation - a high-voltage pulse is used to disrupt the recipient cell membrane to allow the vector through.
- Liposomes - DNA is wrapped in lipid molecules which are fat soluble and thus cross the lipid cell membrane by diffusion.
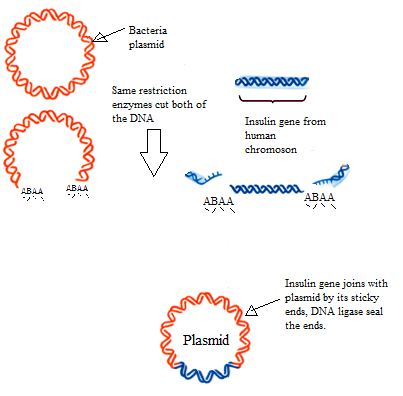
Human insulin
People with type 1 diabetes have an autoimmune disease where their own immune system attacks the insulin producing b-cells in their pancreas. This means that people suffering from the illness cannot produce their own insulin and thus cannot control their blood sugar levels if they get too high. Insulin for clinical use used to be extracted from the pancreas of pigs, however this was fairly inefficient as there was very little insulin in the cells and it also raised a lot of ethical issues regarding the slaughter of the pigs to obtain their insulin. It wasn't until the 1980s where scientists started to use genetically engineered bacteria cells to produce human insulin. This process was a lot more efficient and produced a lot more insulin as well as not requiring the death of pigs!
The steps to produce human insulin are outlined below:
- The mRNA produced from the gene for insulin production was obtained and the enzyme reverse transcriptase was used to synthesise a complementary DNA strand.
- This DNA strand was then put into a PCR (polymerase chain reaction) machine where a second strand of the DNA is produced using free nucleotides and the enzyme DNA polymerase - this produces a copy of the original DNA (i.e. the original gene that codes for insulin) which is called the cDNA.
- Unpaired nucleotides are then added at either end of the DNA strand in order to produce sticky ends that are complementary to the plasmid they will be taken up by.
- The plasmids are then cut using restriction enzyme and mixed with the cDNA gene.
- Some of the plasmids will take up the cDNA and become recombinant plasmids using DNA ligase.
- The plasmids are then mixed with bacteria, some of which will take up the plasmid.
- The bacteria are then grown on an agar plate to produce a colony of clones.
In order to identify which bacteria are transformed or not a process called replica plating is used.
- When the plasmids are cut with restriction enzymes in the process outlined above, the gene for the resistance to the antibiotic tetracycline is also cut using restriction enzymes that have restriction sites in the middle of the tetracycline gene.
- This means that the bacteria that take up the recombinant plasmid will not be resistant to the antibiotic tetracycline but will still be resistant to the antibiotic ampicillin, as that gene has not been cut.
- The bacteria are then grown on agar plates to form colonies.
- Some of the cells from the colonies are transferred onto agar that has been made with ampicillin so that only those that have taken up the recombinant plasmid will grow.
- Other cells from the colonies will be placed onto agar containing the antibiotic tetracycline so that only the non-transformed bacteria (i.e. the bacteria that has not taken up the recombinant plasmid) will grow.
- By keeping track of the different colonies you can identify the transformed bacteria cells which can then be grown on a large scale and the insulin harvested for human use.